_
|
|
|||||||||||||||||||||||
|
High
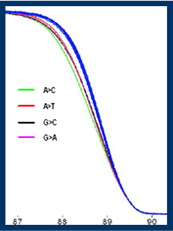
resolution melt analysis
|
||||||||||||||||||||||||
|
Project Leader: Dr Helen White With the development of a new family of saturating DNA dyes, high resolution melt curve analysis (HRM) has been identified as a potentially useful method of high throughput mutation scanning. Recent publications suggest that HRM has a mutation detection sensitivity which is comparable or superior to currently available pre-screening techniques. In 2006, NGRL (Wessex) evaluated 3 machines capable of HRM analysis: RotorGene™ 6000 (Corbett Research), HR-1™ and 384 well LightScanner™ (both from Idaho Technology). Eleven different amplicons were analysed: seven amplicons were generated from the NGRL (Wessex) panel of generic mutation detection control plasmids and 4 were generated from genomic DNA: hMLH1 Exons 1, 7 & 13 and hMSH2 Exon 10. The amplicons varied in size from 139 to 449bp and had GC contents ranging from 22 – 79%. The types of mutations analysed included all possible point mutation base substitutions and 1 and 2bp insertions and deletions. Using the RotorGene™ 6000 (Corbett Research) PCR products were amplified in the presence of the saturating dsDNA binding dye: LC Green® Plus (Idaho Technology). 47 and 70 randomised samples were amplified for each target plasmid and genomic DNA samples respectively (624 samples in total, including controls) and amplification efficiency was monitored in real-time. The same PCR products were analysed using HRM on the RotorGene™ 6000, HR-1™ and 384 well LightScanner™ platforms. Analysis of the RotorGene™ 6000 and HR-1™ melt curves was undertaken manually by two operators and the LightScanner™ data was analysed with the software supplied using high sensitivity settings. Data were then unblinded and the sensitivity and specificity of mutation detection were determined for each amplicon and platform.
|
||||||||||||||||||||||||
Last Updated: 7 August, 2008 by G.Watkins |
||||||||||||||||||||||||